Note
Go to the end to download the full example code.
ASW inference - Three pulses model#
This example implements inference for the ASW population under a three pulses model of admixture, using the tracts package. Inference is performed using autosomal and X chromosome data, allowing for the specification of sex-biased admixture.
To implement this example, we use the following driver file:
samples:
directory: ./TrioPhased/
individual_names: [
"NA19625","NA19700","NA19701","NA19703","NA19704","NA19707","NA19711","NA19712","NA19713","NA19818","NA19819",
"NA19834","NA19835","NA19900","NA19901","NA19904","NA19908","NA19909","NA19913","NA19914","NA19916","NA19917",
"NA19920","NA19921","NA19922","NA19923","NA19982","NA19984","NA20126","NA20127","NA20274","NA20276","NA20278",
"NA20281","NA20282","NA20287","NA20289","NA20291","NA20294","NA20296","NA20298","NA20299","NA20314","NA20317",
"NA20318","NA20320","NA20321","NA20332","NA20334","NA20339","NA20340","NA20342","NA20346","NA20348","NA20351",
"NA20355","NA20356","NA20357","NA20359","NA20362","NA20412"]
male_names : [
"NA19700","NA19703","NA19711","NA19818","NA19834","NA19900","NA19904","NA19908","NA19916","NA19920",
"NA19922","NA19982","NA19984","NA20126","NA20278","NA20281","NA20291","NA20298","NA20318","NA20340",
"NA20342","NA20346","NA20348","NA20351","NA20356","NA20362"] #see Readme_dataprocessing.md for how this was generated
filename_format: "{name}_{label}_final.bed"
labels: [A, B] #If this field is omitted, 'A' and 'B' will be used by default
chromosomes: 1-22 #The chromosomes to use for analysis. Can be specified as a list or a range
allosomes: [X]
output_filename_format: "ASW_test_output_{label}"
log_filename: 'ASW_three_pulses.log'
output_directory: ./output_three_pulses/
verbose_log: 1
verbose_screen: 30
log_scale : True
start_params:
t1: 10
REUR: 0.8
RAFR: 0.9
REUR2: 0.2
t2: 5
t3: 3
REUR_sex_bias: 0.1
REUR2_sex_bias: 0.1
RAFR_sex_bias: 0.1
repetitions: 3
seed: 100
maximum_iterations: 1000
unknown_labels_for_smoothing: ["UNK", "centromere","miscall"] # segments with these labels will be smoother over, that is, will be filled with neighbouring ancestries up to their midpoints.
exclude_tracts_below_cm: 2
npts : 50
#fix_parameters_from_ancestry_proportions: ['REUR2', 'RAFR', 'REUR2_sex_bias', 'RAFR_sex_bias']
output_directory: ./output_three_pulses/
ad_model_autosomes: M
ad_model_allosomes: DC
Complete results from this analysis are saved in the output directory specified in the driver file. Below, we display the optimal parameters estimated from this analysis, as well as the plots illustrating the inferred tract length distributions, compared to the observed histograms, for every source population and chromosome type (autosomes and X chromosome).
Optimal parameters#
parameter |
value |
|---|---|
REUR |
0.7729121375993303 |
REUR_sex_bias |
0.1132088204931403 |
t1 |
8.634748344515474 |
RAFR |
0.8841928552747739 |
RAFR_sex_bias |
-0.004658383791999676 |
t2 |
7.226408784836712 |
REUR2 |
0.14493237980731358 |
REUR2_sex_bias |
-0.435333740753701 |
t3 |
5.666278423687627 |
likelihood -978.508 |
Tract length histograms#
Autosomal admixture#
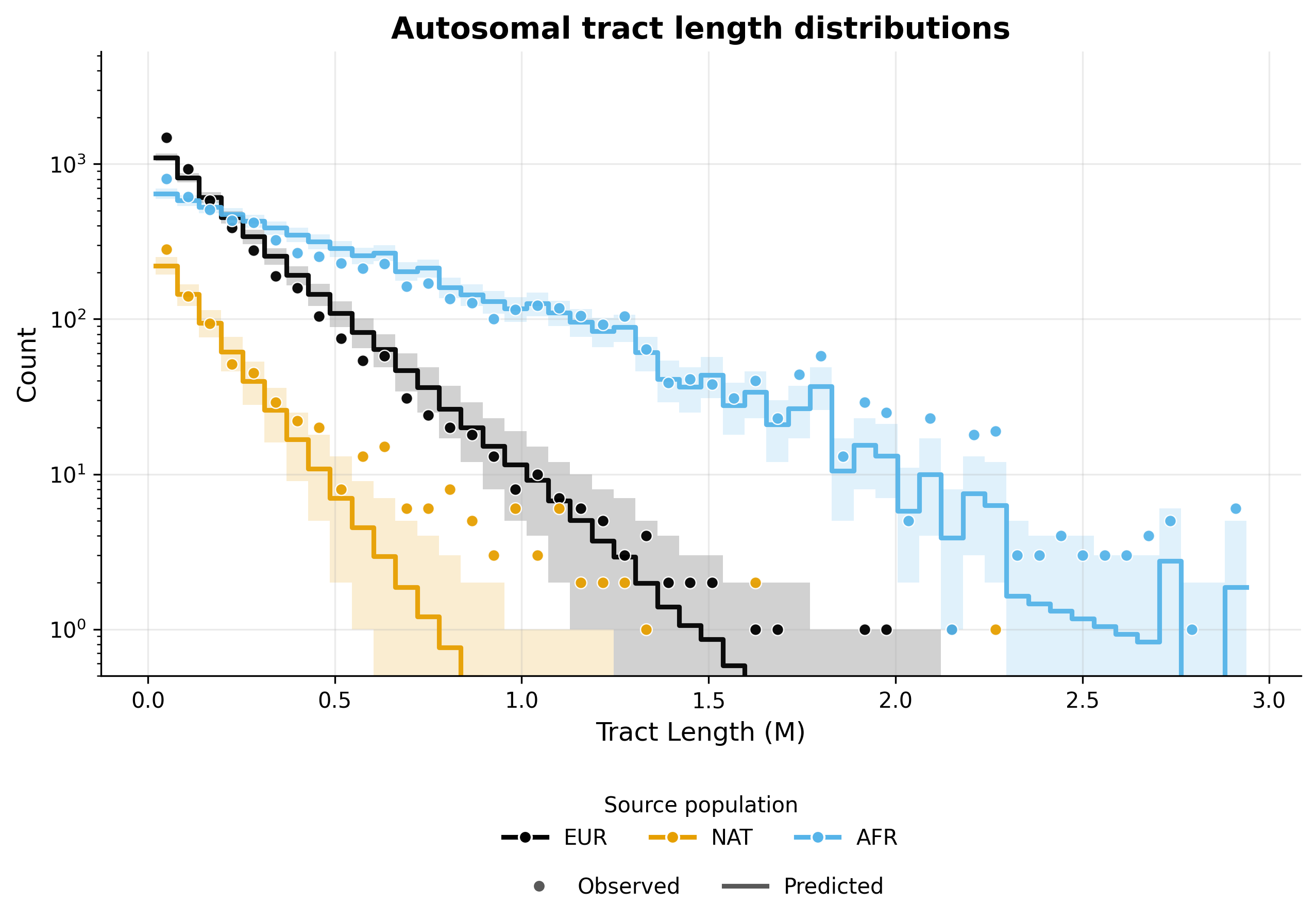
X chromosome admixture in females#
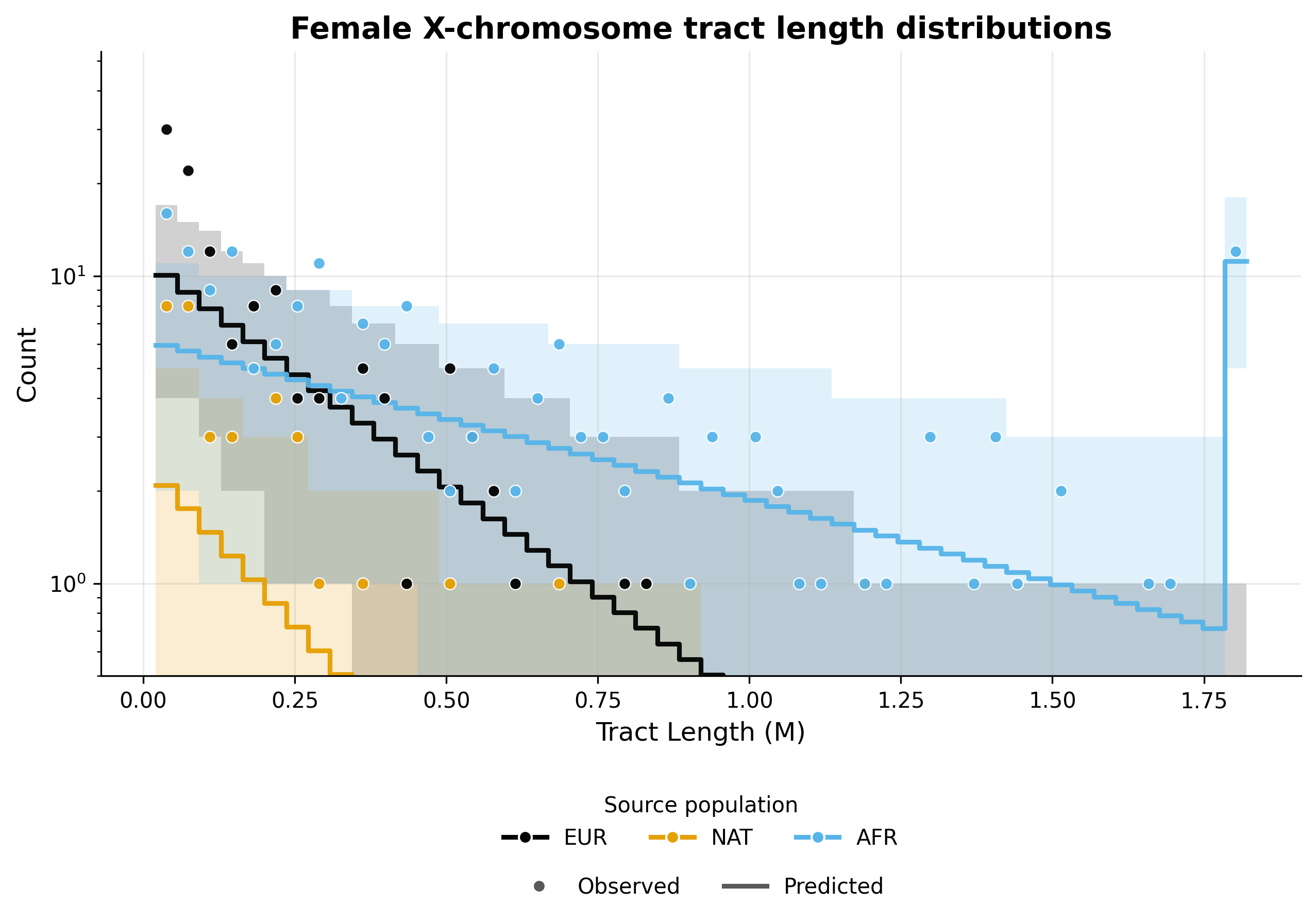
X chromosome admixture in males#
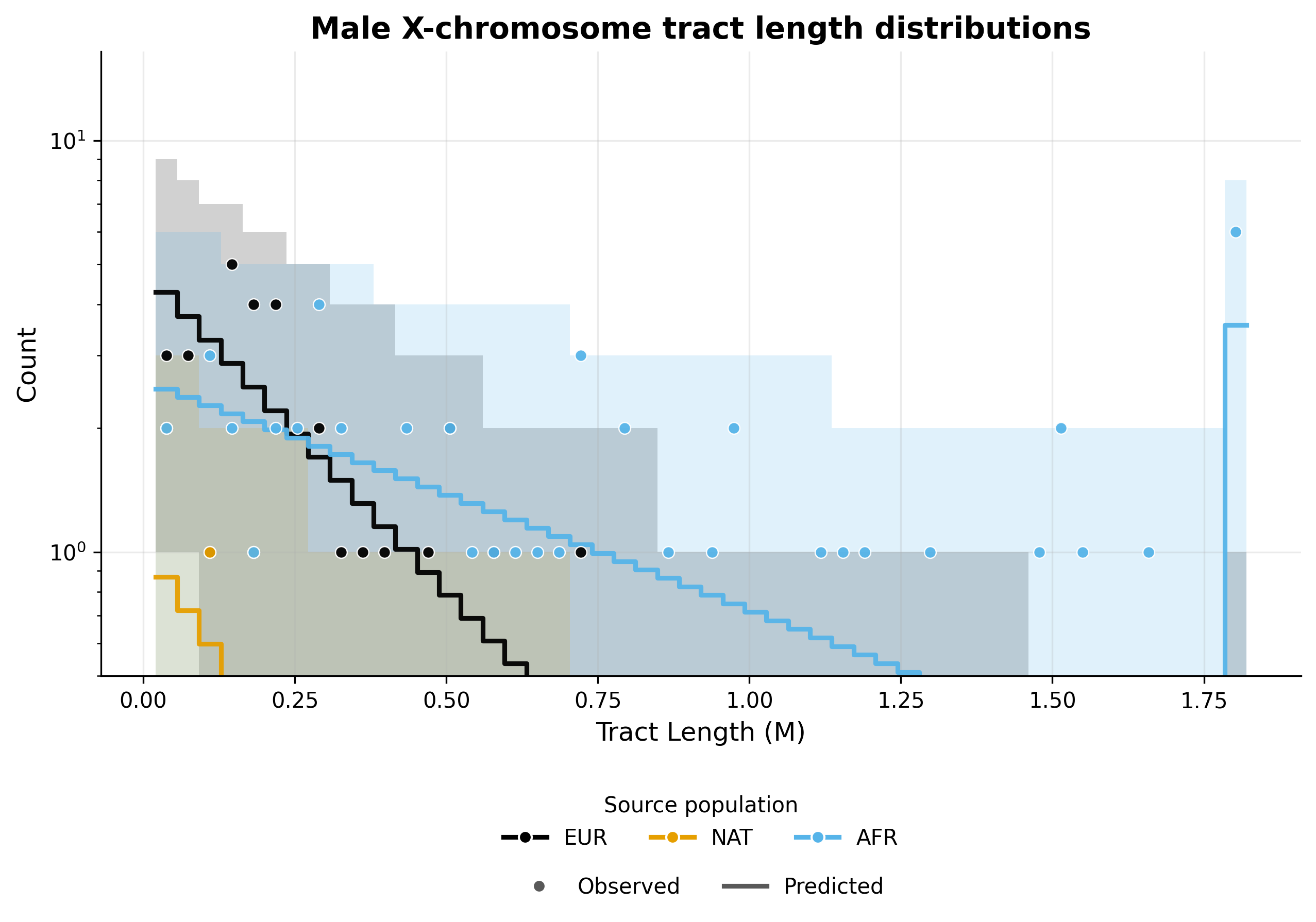
------------------------------------------------------------------------------------------------
Running tracts 2.0 with driver file: ASW_three_pulses.yaml
------------------------------------------------------------------------------------------------
Results will be written to: output_three_pulses.
Using log file: output_three_pulses/ASW_three_pulses.log.
excluding_tracts_below set to 2.0 cM.
Ancestries: EUR, NAT, AFR
Data autosome proportions: [0.19578862 0.03825495 0.76595643]
Data allosome proportions: [0.16839124 0.03818939 0.79341937]
Model parameters: REUR, REUR_sex_bias, t1, RAFR, RAFR_sex_bias, t2, REUR2, REUR2_sex_bias, t3
Admixture is modelled with the M model for autosomes and with the DC model for allosomes.
Multiple starting parameters will be generated and used for multiple optimization runs.
-----------------------------------------------------------------------------------------------------
Step 1 : Optimizing autosomal likelihood over parameters ['REUR', 't1', 'RAFR', 't2', 'REUR2', 't3'].
-----------------------------------------------------------------------------------------------------
Starting parameters for step 1 optimization
---------------------------------------------------------------------------------------------
Run | REUR | t1 | RAFR | t2 | REUR2 | t3
---------------------------------------------------------------------------------------------
1 | 0.8 | 9.867 | 0.9 | 6.789 | 0.2 | 3.064
2 | 0.8 | 9.854 | 0.9 | 6.8 | 0.2 | 2.506
3 | 0.8 | 11.99 | 0.9 | 5.966 | 0.2 | 3.074
---------------------------------------------------------------------------------------------
Optimization run #1
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
30 , -1215.63 , array([ 0.797778 , 0 , 9.96533 , 0.891678 , 0 , 6.94546 , 0.198058 , 0 , 3.68714 ]), Autosomes
60 , -889.862 , array([ 0.796526 , 0 , 9.58264 , 0.885576 , 0 , 7.3138 , 0.190549 , 0 , 4.30489 ]), Autosomes
90 , -763.181 , array([ 0.786673 , 0 , 9.03934 , 0.880502 , 0 , 7.57498 , 0.166511 , 0 , 4.80616 ]), Autosomes
120 , -727.807 , array([ 0.786846 , 0 , 8.85361 , 0.878754 , 0 , 7.64575 , 0.15868 , 0 , 4.89178 ]), Autosomes
150 , -714.745 , array([ 0.785267 , 0 , 8.84535 , 0.878385 , 0 , 7.67761 , 0.154349 , 0 , 4.94699 ]), Autosomes
180 , -703.543 , array([ 0.783706 , 0 , 8.8365 , 0.877875 , 0 , 7.68805 , 0.150951 , 0 , 4.99474 ]), Autosomes
Optimization completed
----------------------
Optimization run #2
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
30 , -1536.14 , array([ 0.798213 , 0 , 10.1199 , 0.8867 , 0 , 6.93854 , 0.197798 , 0 , 2.91645 ]), Autosomes
60 , -983.654 , array([ 0.797388 , 0 , 10.0798 , 0.874671 , 0 , 7.69938 , 0.18867 , 0 , 3.3935 ]), Autosomes
90 , -811.131 , array([ 0.79505 , 0 , 9.23515 , 0.863564 , 0 , 7.95468 , 0.166934 , 0 , 3.96751 ]), Autosomes
120 , -779.995 , array([ 0.79503 , 0 , 9.17526 , 0.863572 , 0 , 8.03091 , 0.161757 , 0 , 4.02618 ]), Autosomes
Optimization completed
----------------------
Optimization run #3
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
30 , -1344.44 , array([ 0.797258 , 0 , 12.0966 , 0.893864 , 0 , 6.61638 , 0.198304 , 0 , 3.49862 ]), Autosomes
60 , -995.334 , array([ 0.792664 , 0 , 11.3806 , 0.882393 , 0 , 6.81238 , 0.188567 , 0 , 4.27351 ]), Autosomes
90 , -821.322 , array([ 0.782795 , 0 , 10.2064 , 0.878202 , 0 , 6.96528 , 0.172819 , 0 , 5.07204 ]), Autosomes
120 , -737.571 , array([ 0.780019 , 0 , 9.62516 , 0.8825 , 0 , 7.1452 , 0.158982 , 0 , 5.38853 ]), Autosomes
150 , -673.49 , array([ 0.774047 , 0 , 8.65556 , 0.884256 , 0 , 7.20062 , 0.147654 , 0 , 5.66046 ]), Autosomes
180 , -667.882 , array([ 0.773252 , 0 , 8.63872 , 0.884272 , 0 , 7.23006 , 0.145465 , 0 , 5.66193 ]), Autosomes
210 , -666.834 , array([ 0.772967 , 0 , 8.63523 , 0.884189 , 0 , 7.2254 , 0.145051 , 0 , 5.66488 ]), Autosomes
Optimization completed
----------------------
Step 1: Results from multiple optimization runs with different starting parameters:
-------------------------------------------------------------------------------------------------------------------------------------------------------------
Run | LogLik | REUR | REUR_sex_bias | t1 | RAFR | RAFR_sex_bias | t2 | REUR2 | REUR2_sex_bias | t3
-------------------------------------------------------------------------------------------------------------------------------------------------------------
1 | -703.156 | 0.7836 | 0 | 8.837 | 0.8778 | 0 | 7.687 | 0.1509 | 0 | 4.993
2 | -779.341 | 0.795 | 0 | 9.177 | 0.8636 | 0 | 8.033 | 0.1617 | 0 | 4.026
3 | -666.538 | 0.7729 | 0 | 8.635 | 0.8842 | 0 | 7.226 | 0.1449 | 0 | 5.666
-------------------------------------------------------------------------------------------------------------------------------------------------------------
Selecting best parameters from step 1 and proceeding to step 2 optimization.
----------------------------------------------------------------------------------------------------------------
Step 2 : Optimizing allosomal likelihood over parameters : ['REUR_sex_bias', 'RAFR_sex_bias', 'REUR2_sex_bias'].
----------------------------------------------------------------------------------------------------------------
Starting parameters for step 2 optimization (non-sex-bias parameters are fixed to the best step 1 estimates).
----------------------------------------------------------------------------------------------------------------------------------------------
Run | REUR | REUR_sex_bias | t1 | RAFR | RAFR_sex_bias | t2 | REUR2 | REUR2_sex_bias | t3
----------------------------------------------------------------------------------------------------------------------------------------------
1 | 0.7729 | 0.4249 | 8.635 | 0.8842 | 0.004503 | 7.226 | 0.1449 | -0.2643 | 5.666
2 | 0.7729 | 0.1154 | 8.635 | 0.8842 | -0.0003643 | 7.226 | 0.1449 | -0.434 | 5.666
3 | 0.7729 | 0.08855 | 8.635 | 0.8842 | 0.00242 | 7.226 | 0.1449 | 0.2594 | 5.666
----------------------------------------------------------------------------------------------------------------------------------------------
Optimization run #1
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
30 , -216.292 , array([ 0.772912 , 0.424785 , 8.63475 , 0.884193 , -0.00464054 , 7.22641 , 0.144932 , -0.265427 , 5.66628 ]), Female allosomes
30 , -95.5145 , array([ 0.772912 , 0.424785 , 8.63475 , 0.884193 , -0.00464054 , 7.22641 , 0.144932 , -0.265427 , 5.66628 ]), Male allosomes
Optimization completed.
-----------------------
Optimization run #2
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
Optimization completed.
-----------------------
30 , -668.561 , array([ 0.772912 , 0.113209 , 8.63475 , 0.884193 , -0.00465838 , 7.22641 , 0.144932 , -0.435334 , 5.66628 ]), Autosomes
30 , -214.665 , array([ 0.772912 , 0.113209 , 8.63475 , 0.884193 , -0.00465838 , 7.22641 , 0.144932 , -0.435334 , 5.66628 ]), Female allosomes
30 , -95.282 , array([ 0.772912 , 0.113209 , 8.63475 , 0.884193 , -0.00465838 , 7.22641 , 0.144932 , -0.435334 , 5.66628 ]), Male allosomes
Optimization run #3
Iter. Log-likelihood Model parameters Transmission
-------------------------------------------------------------
30 , 2.67695e+26 , array([ 0.772912 , 0.0875555 , 8.63475 , 0.884193 , -0.00469233 , 7.22641 , 0.144932 , 0.257968 , 5.66628 ]), OOB (oob=-2.6769533421067138e-06)
Optimization completed.
-----------------------
Step 2: Results from multiple optimization runs with different starting parameters:
-------------------------------------------------------------------------------------------------------------------------------------------------------------
Run | LogLik | REUR | REUR_sex_bias | t1 | RAFR | RAFR_sex_bias | t2 | REUR2 | REUR2_sex_bias | t3
-------------------------------------------------------------------------------------------------------------------------------------------------------------
1 | -311.806 | 0.7729 | 0.4248 | 8.635 | 0.8842 | -0.004668 | 7.226 | 0.1449 | -0.2654 | 5.666
2 | -309.947 | 0.7729 | 0.1132 | 8.635 | 0.8842 | -0.004658 | 7.226 | 0.1449 | -0.4353 | 5.666
3 | -310.182 | 0.7729 | 0.08757 | 8.635 | 0.8842 | -0.004644 | 7.226 | 0.1449 | 0.258 | 5.666
-------------------------------------------------------------------------------------------------------------------------------------------------------------
Selecting best parameters from step 2.
Step 2 used allosomal data only. Final likelihood is evaluated on autosomal + allosomal data at the selected optimal parameters.
Final parameters and corresponding likelihood computed on autosomal + allosomal data:
-------------------------------------------------------------------------------------------------------------------------------------------------------
LogLik | REUR | REUR_sex_bias | t1 | RAFR | RAFR_sex_bias | t2 | REUR2 | REUR2_sex_bias | t3
-------------------------------------------------------------------------------------------------------------------------------------------------------
-978.508 | 0.7729 | 0.1132 | 8.635 | 0.8842 | -0.004658 | 7.226 | 0.1449 | -0.4353 | 5.666
-------------------------------------------------------------------------------------------------------------------------------------------------------
Results saved to : output_three_pulses
{'destination_dir': PosixPath('/home/runner/work/tracts/tracts/docs/source/auto_examples/ASW/output_three_pulses'), 'table_file': PosixPath('/home/runner/work/tracts/tracts/docs/source/auto_examples/ASW/output_three_pulses/ASW_test_output_optimal_parameters.txt')}
import sys
from pathlib import Path
sys.path.append('.')
from tracts.driver import run_tracts
script_path = Path(sys.argv[0]).resolve()
driver_filename = "ASW_three_pulses.yaml"
run_tracts(driver_filename = driver_filename, script_dir = script_path)
# Don't run the code below: for documentation purposes only.
from tracts.doc_utils import prepare_example_outputs_for_docs
prepare_example_outputs_for_docs("output_three_pulses")
Total running time of the script: (1 minutes 49.460 seconds)